PCR and antibody-based tests for SARS-CoV-2 infection are a key part of the efforts to combat the COVID-19 pandemic. But as this case report shows, PCR testing for SARS-CoV-2 can sometimes provide more questions than answers.
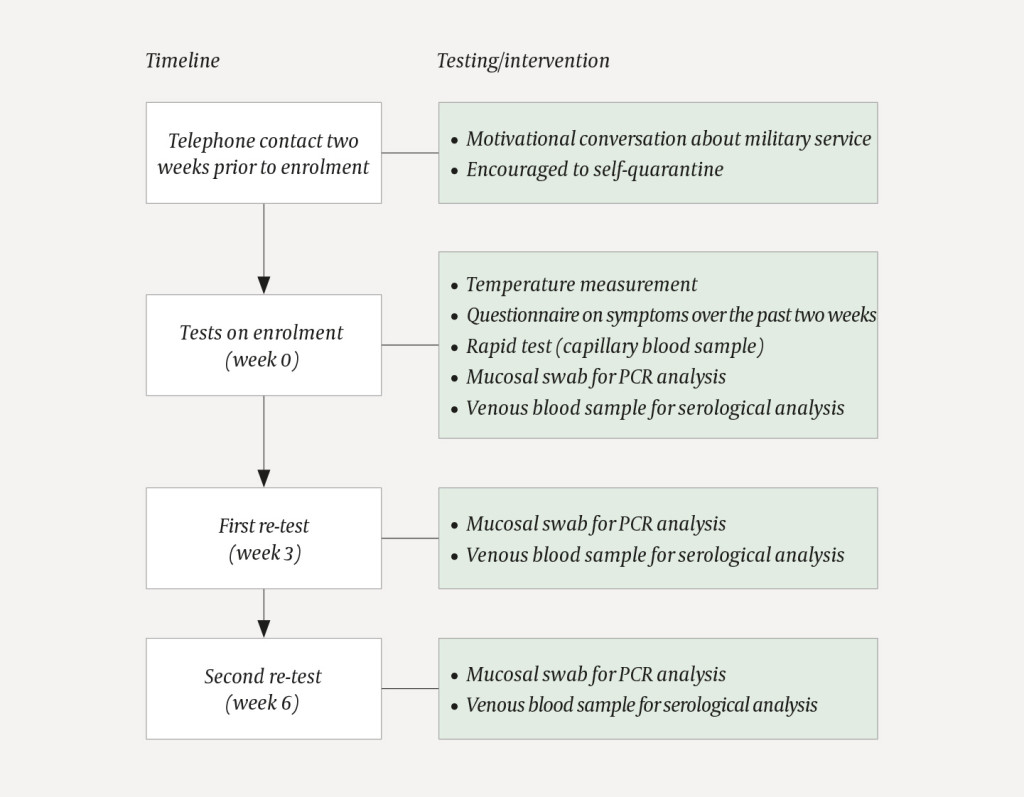
A previously healthy man responded to the challenges posed by the COVID-19 pandemic by voluntarily self-isolating from March 2020 until the following month, when he was due to begin military service at the KNM Harald Haarfagre basic training centre at Madla in Stavanger. While at home, he had sole use of one of the house's two bathrooms, but he ate meals together with other members of the household. In mid-March, the household received a visit from the prospective recruit's sister, who had mild cold-like symptoms, but not to the extent that anyone suspected COVID-19. No-one else in the family had had respiratory symptoms or a suspected coronavirus infection, but neither had they undergone any form of testing. In connection with the arrival of new recruits for the induction training programme, the Norwegian Armed Forces had put in place a system of extended systematic infection screening (1). This consisted of a detailed personal health statement and temperature measurement upon arrival, as well as a rapid test to detect antibodies against SARS-CoV-2, a venous blood sample for serological analysis of antibodies, and a nasopharyngeal swab to detect SARS-CoV-2 RNA in the upper respiratory tract (by means of polymerase chain reaction, PCR). PCR and serological tests were repeated as a matter of routine for all recruits after three and six weeks of the eight-week induction training programme (Figure 1).
Upon enrolment, the man was afebrile with no respiratory symptoms. However, he tested positive for IgG and negative for IgM in an anti-SARS-CoV-2 rapid antibody test. He was placed in quarantine, but this was revoked the next day when his PCR result proved to be negative. He entered the standard induction training programme and then continued with routine testing. Serological analysis of a specimen collected on enrolment was also positive (Table 1).
Table 1
Patient's test results for SARS-CoV-2
| Test |
Week 0 |
Week 3 |
Week 6 |
Week 6 |
|---|---|---|---|---|
| Rapid test1 |
Positive (IgG) |
- |
- |
- |
| PCR test2 |
Negative |
Negative |
Positive |
Negative (× 2) |
| Serological test4 (S/Co)5 |
Positive (18.7) |
Positive (20.3) |
Positive (23.2) |
- |
1Acro 2019-nCoV IgG/IgM rapid antibody test
2Cobas SARS-CoV-2
3Ct =cycle threshold.
4Elecsys IgM/IgG Anti-SARS-CoV-2 assay
5The analysis yields a total measure of IgG and IgM, expressed as an index which is the ratio of the signal measured in the sample (IgG and IgM) to the signal from an internal control in the assay. The manufacturer refers to this as the signal-to-cut-off ratio, abbreviated to S/Co.
Three weeks after enrolment, the recruit still had no symptoms, and a routine follow-up PCR test was negative. However, routine serological testing revealed rising antibody levels. At week 6, he was still asymptomatic, but a routine nasopharyngeal swab tested positive on PCR. The swab was analysed by the same laboratory using the same method as for the two previous PCR tests. The serological assays also showed a further increase in the recruit's antibody levels (Table 1).
The positive PCR test was reported to the chief municipal medical officer and the Norwegian Surveillance System for Communicable Diseases, and the recruit was placed in isolation. The next day, in consultation with the healthcare authorities, control specimens were collected and were analysed at two different laboratories. Both controls tested negative (Table 1). After consultation with the Norwegian Institute of Public Health and the chief infection control medical officer in Stavanger municipality, the recruit was taken out of isolation. He completed the remainder of the induction training programme without developing any symptoms, and has not been ill since finishing the programme in June 2020.
Discussion
The soldier had no COVID-19-associated symptoms before, during or after his stay at the training centre. Nevertheless, he had a positive rapid antibody test on enrolment, positive serological assays with slightly increasing anti-SARS-CoV-2 antibody levels, and a weakly positive PCR test for SARS-CoV-2 RNA at the last testing session in week 6.
The rising anti-SARS-CoV-2 antibody levels in our patient are difficult to interpret. IgG/IgM antibodies are reported to develop between a few days and possibly up to a week after symptom onset in cases of COVID-19. The time course is somewhat different for IgG and IgM, but the concentration reaches a maximum several weeks after the patient first had symptoms, partly as a result of antibody avidity (2, 3). The serodynamics in asymptomatic patients have yet to be fully characterised.
The patient's positive serology results for SARS-CoV-2 may indicate a history of infection prior to arrival at the training centre, which was not detected in the anamnesis or via the first PCR tests. The test used (Roche Elecsys Anti-SARS-CoV-2) has a sensitivity of more than 95 % and a specificity of more than 99 % (4).
Non-specific reactivity or cross-reactivity with antibodies against other coronaviruses may have led to false positive serological results. However, the S/Co (signal-to-cut-off ratio) would most likely have been lower if that were the case, and would not have shown successive increases over the brief period in which antibody levels were measured.
The weakly positive PCR result for SARS-CoV-2 RNA in the last routine test in week 6 could not be reproduced. This might mean that the result was a false positive, or it could reflect the presence of non-viable virus or of very small amounts of viral RNA (5).
The results of the PCR tests for the first two samples and the control samples might have been false negatives. Other have also reported cases of a positive PCR test after two previous negative tests 24 hours apart, and have proposed that this may be due to false negative results, reinfection or reactivation (6). Another possible explanation is that the virus was strongly bound to receptors in the lower respiratory tract and was not expelled as the patient had no cough. Antibody coating of the virus can also make it difficult to detect antigens, but will not affect the detection of RNA. Furthermore, shedding of the SARS-CoV-2 virus has yet to be fully characterised, especially late in the disease course (7). A mix-up of samples is a theoretical possibility, but is highly unlikely given the stringent procedures followed.
Overall, we believe that the results of the serological assays and PCR tests together suggest a previous asymptomatic infection prior to arrival at the training centre.
The fact that our patient did not infect others may suggest low transmissibility. However, it is not a given that patients with asymptomatic infection will be less likely to transmit the virus than symptomatic patients. The high Ct (cycle threshold) value of over 35 in the positive PCR test indicates that there must have been little viral RNA present in the sample. Another study has shown that SARS-CoV-2 isolated from mucosal specimens with high PCR-Ct values cannot be cultured and are therefore no longer viable (8). The absence of infection transmission may also reflect the effects of good infection control measures or may indicate that the patient was not infectious.
This case report demonstrates that a repeat PCR test can detect viral RNA even if previous tests have been negative and there has been no intercurrent disease. It also shows concordance between positive results for antibody-based rapid tests and tests on venous blood samples, and illustrates how serological assays can be helpful when interpreting PCR results. Finally, this case highlights the various challenges that can arise when screening asymptomatic individuals on a large scale.
The patient has consented to the publication of this article.
The article has been peer-reviewed.
- 1.
Norheim AJ, Nakstad E, Berg AS et al. Testing Armed Forces recruits for COVID-19. Tidsskr Nor Legeforen 2020; 140. doi: 10.4045/ tidsskr.20.0384. [PubMed][CrossRef]
- 2.
Lijia S, Lihong S, Huabin W et al. Serological chemiluminescence immunoassay for the diagnosis of SARS-CoV-2 infection. J Clin Lab Anal 2020; 34: e23466. [PubMed][CrossRef]
- 3.
Xiang F, Wang X, He X et al. Antibody Detection and Dynamic Characteristics in Patients With Coronavirus Disease 2019. Clin Infect Dis 2020; 71: 1930–4. [PubMed][CrossRef]
- 4.
Evaluation of sensitivity and specificity of four commercially available SARS-CoV-2 antibody immunoassays London: Public Health England, University of Oxford, Oxford University Hospitals NHS Foundation Trust, 2020. https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/898437/Evaluation__of_sensitivity_and_specificity_of_4_commercially_available_SARS-CoV-2_antibody_immunoassays.pdf Accessed 2.11.2020.
- 5.
Wölfel R, Corman VM, Guggemos W et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020; 581: 465–9. [PubMed][CrossRef]
- 6.
Sethuraman N, Jeremiah SS, Ryo A. Interpreting diagnostic tests for SARS-CoV-2. JAMA 2020; 323: 2249–51. [PubMed][CrossRef]
- 7.
Yan D, Liu XY, Zhu YN et al. Factors associated with prolonged viral shedding and impact of lopinavir/ritonavir treatment in hospitalised non-critically ill patients with SARS-CoV-2 infection. Eur Respir J 2020; 56: 2000799. [PubMed][CrossRef]
- 8.
La Scola B, Le Bideau M, Andreani J et al. Viral RNA load as determined by cell culture as a management tool for discharge of SARS-CoV-2 patients from infectious disease wards. Eur J Clin Microbiol Infect Dis 2020; 39: 1059–61. [PubMed][CrossRef]
Dersom rekrutten hadde ervervet immunitet ved tidligere gjennomgått infeksjon så forklarer dette prøvesvar ved ankomst til tjeneste. Dersom han ble utsatt for en lav mengde covid-19-smitte under oppholdet så kan dette kanskje forklare en lett stigning i antistoff og en svakt positiv PCR og senere negative PCR (etter at antistoffene og immunsystemet har eliminert eller «coatet» den nye smitten).